In the largest-ever collaborative study of its kind, a team of cardiovascular genetics researchers from around the world have identified 13 new genetic variants associated with coronary artery disease (CAD). This finding more than doubled the number of genetic variants known to impact risk for CAD. The study also confirmed the association of 10 previously identified variants.
“If any individual has a large combination of these variants, their risk is about four times that of somebody who has the fewest number. That’s one of the most important elements of our discovery.” – Alexandre Stewart, Principal Investigator, Ruddy Canadian Cardiovascular Genetics Centre, UOHI
Of the 23 genes found or verified, only six were found to be related to traditional risk factors, such as blood cholesterol levels. “This is a landmark study and the implications are twofold,” said Dr. Robert Roberts, President and CEO of the University of Ottawa Heart Institute. “One is that there’s a whole group of undiscovered mechanisms behind the development of coronary artery disease that is a possible target for drug development. The second is that we now have a fair number of genes to screen for in terms of determining risk in the setting of primary prevention.”
The collaborative study, published in Nature Genetics, is called CARDIoGRAM (for Coronary Artery Disease Genome-Wide Replication and Meta-Analysis). Two additional studies published in the same issue, from a separate collaboration, called the Coronary Artery Disease (C4D) Genetics Consortium and from the Huazhong University of Science & Technology in China, found six additional variants associated with heart disease, bringing the total number of genetic risk factors discovered through genome-wide association studies (GWAS) to slightly less than 30.
More Patients, Stronger Results
Led by the University of Lübeck in Germany, the CARDIoGRAM effort included researchers from more than 100 organizations, including the Ruddy Canadian Cardiovascular Genetics Centre at the Heart Institute.
From inception to results, the collaborators moved at breakneck speed to finish the study. “We all had reached the same impasse in our research. To make any new discoveries in this field, we needed to bring our data together,” explained Alexandre Stewart, Principal Investigator at the Ruddy Canadian Cardiovascular Genetics Centre and Head of the Heart Institute’s Genetics Core Laboratory.
CARDIoGRAM was a meta-analysis, a type of study that collectively analyzes data from many other studies. Fourteen previous GWAS, including 22,233 patients diagnosed with CAD and 64,762 control patients, all of European descent, were included in the CARDIoGRAM analysis.
GWAS search for tiny variations in DNA—called single-nucleotide polymorphisms (SNPs)—that are more likely to be found in people with a disease than in people without that disease. “Most of these variations have small effects on risk, and so to identify a genetic marker that increases risk by, say, 10 per cent, you really need very large sample sizes,” explained Dr. Ruth McPherson, Heart Institute physician-scientist and Director of the Atherogenomics Laboratory, who spearheaded the Ruddy Centre’s participation in CARDIoGRAM.
To complicate the genetic risk calculations in heart disease, 85 per cent of the population older than 50 years of age is thought to have some degree of measurable CAD, explained Stewart, meaning that the differences in disease burden in patients and so-called controls is likely to be relatively small.
In addition, said Stewart, “with individual studies, even if you have a large sample size, it’s still one experiment in one regional population. For GWAS results to have meaning globally, you need to bring everybody together, and that’s what we did.”
The Long Road from Bench to Bedside
More than three years out from the discovery of the 9p21 risk variant, scientists are gathering the first solid clues as to how genetic variations in that region of the genome influence the risk of heart disease.
The 9p21 variant is found in a so-called gene desert—an area of the genome containing no known genes or functional nucleotides. This puzzled scientists, as 9p21 showed such a strong association with heart disease.
But non-coding areas of the genome like 9p21 can still influence the expression of nearby genes. In 2009, Heart Institute researchers found the first clue as to how 9p21 might alter cell functioning. They found that changes in 9p21 alter the function of a regulatory gene called ANRIL, which in turn affects the expression of other genes linked to cell division and atherosclerosis.
This year, scientists at the University of California, San Diego, found another piece of the puzzle, with research results suggesting that people with genetic variations in 9p21 may have an altered response to inflammation, the body’s natural response to injury and invasion.
The researchers followed an intricate molecular pathway from 9p21 to its effects on a type of DNA sequence called an enhancer, which may serve to regulate an immune system protein called interferon-gamma. That protein can control other genes involved in the inflammatory process.
Whether ANRIL and this newly discovered immune pathway are related or separately influenced by 9p21 is not known. “I think they’re working together, but the biology will be found to be quite complicated,” commented Alexandre Stewart, Principal Investigator at the Ruddy Canadian Cardiovascular Genetics Centre.
Research published in late 2010 by Heart Institute researchers in the Journal of the American College of Cardiology provides important new information about the end result of 9p21’s influence on heart disease. Using measurements of arterial blockage from 950 patients with early-onset heart disease and another group of 764 patients with late-onset disease, the researchers found that patients with two copies of a variation in 9p21 were more likely than those with one or no copies to have severe plaque deposits in several blood vessels. Patients with one copy of the risk variant were more likely than those with no variations to have severe plaque deposits in several blood vessels.
“If you have two copies of the risk variant, you’re at much higher risk of having more severe disease,” stressed Stewart. In contrast, the number of altered copies of 9p21 showed no association with the risk for a heart attack. This indicates that the cellular effects of the 9p21 risk variant lead to the deposit of coronary plaques but are not the direct causes of heart attack which can result from the rupture of existing plaques or the formation of blood clots.
Collective Impact
After the main CARDIoGRAM analysis was performed, the results were confirmed in 56,683 additional samples—about half patients with CAD and half controls—bringing the total number of people studied to more than 140,000.
“When we say this panel of variants accounts for 10 per cent of the population-based risk, it’s important to realize that that’s not the individualized risk,” explained Stewart. “If any individual has a large combination of these risk variants, their risk is about four times that of somebody who has the fewest number. That’s one of the most important elements of our discovery—if you look at the extremes of genotypes, the ones with the most risk variants and the ones with the least, you see this fourfold difference at a personal level. These variants can have a huge impact on individuals.”
An Expanding Map of Genetic Risk
A total of 29 genetic variations have now been found to be associated with heart disease. However, the work of understanding how these small changes in the genetic code affect risk is just beginning. Most of these variants are not associated with any known risk factors for heart disease. Some have been shown to alter the risk of other common diseases, but the functional pathways of most remain unknown.
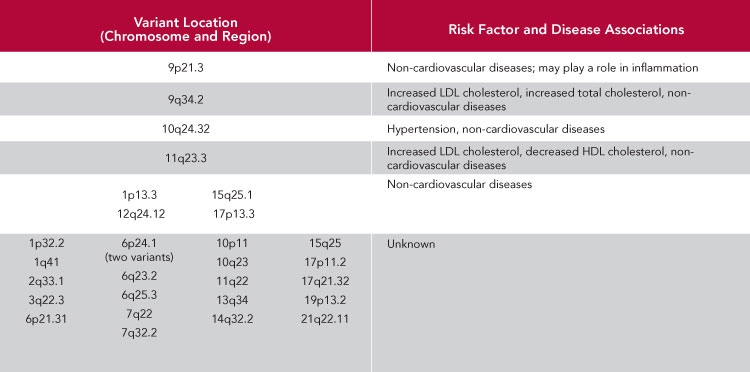
What Those Numbers and Letters Mean
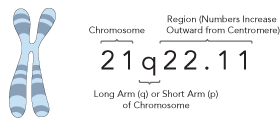
The location of a genetic variation within the genome is written as: 21q22.11. The first number, in this case 21, refers to which of the 23 pairs of human chromosomes the variant is found on. All chromosomes have a short arm (p) and a long (q) arm. The numbers that follow refer to the regions of a chromosome in much the way that a street address specifies a particular house. Numbering starts at the centromere—the point at which the arms of a chromosome are joined—and increases towards the end of the arm.
In addition, commented Dr. McPherson, these findings point to larger inter-connected molecular pathways that may promote the development of heart disease. Other genetic variations along these pathways—including rarer higher risk variants—may also be targets for future research.
Interestingly, five of the newly identified genes and three of those previously confirmed were also strongly associated with the risk of other diseases as varied as celiac disease, diabetes and lung cancer. Further studies are needed to understand whether these associations indicate a common link in disease origin, or whether multiple genes in an identified region of DNA are independently contributing to different diseases, explained the authors.
Preliminary studies have shown that several of the genetic variants identified earlier in Caucasian patients also affect risk in Asian and South Asian populations, though these early results need to be confirmed in larger studies. Other studies are actively looking for genetic risk factors in populations of African decent, which have been proven to have greater genetic variation between individuals, making GWAS even more challenging.
As for the population covered by CARDIoGRAM, “there will be more to discover,” commented Stewart. The researchers are looking at an even larger collaboration with the C4D investigators, involving patients of both European and Asian descent.
But the bigger picture remains the same, said Dr. Roberts. “Now our job is to understand how these genes work, develop a new group of drugs to target them and identify people who will benefit most to reduce their risk of heart attack and other cardiac events.”

